
Pharyngeal microbiota was mainly composed of Streptococcus, Rothia, etc. The intestinal microbiota was mainly composed of Unidentified Clostridiales, Klebsiella, Unidentified Enterobacteriaceae, Enterobacter, Streptococcus, etc. At T28, there was a significant difference in microbial composition between intestine and pharynx ( p < 0.001). Ureaplasma and Fusobacterium were detected in both intestine and pharynx. The pharyngeal microbiota was mainly composed of Ureaplasma, Bacteroides, Fusobacterium, etc. The intestinal microbiota was mainly composed of Unidentified Enterobacteriaceae, Ralstonia, Streptococcus, Fusobacterium, Ureaplasma, etc. Results: At T1, the difference in microbial composition between intestine and pharynx was not statistically significant. Based on the sequencing results, the composition of the intestinal and pharyngeal microbiota was compared and analyzed. 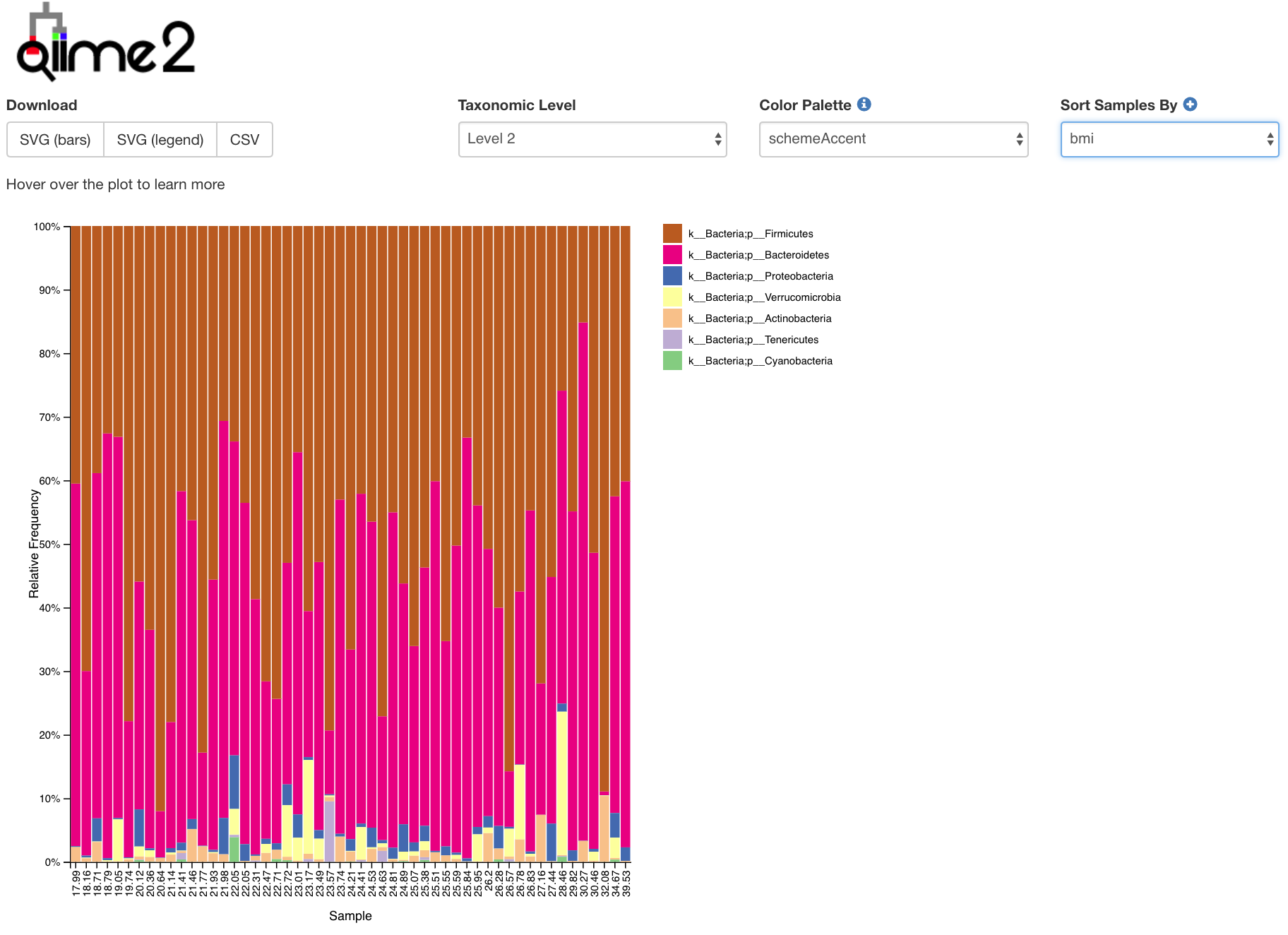
Total bacterial DNA was extracted and sequenced using the Illumina MiSeq Sequencing System based on the V3–V4 hyper-variable regions of the 16S rRNA gene. Stool samples and pharyngeal swabs samples were collected from each neonate on the first day (T1) and the 28th day (T28) after birth. Methods: Thirteen neonates born at 26–32 weeks gestational age (GA) hospitalized at the neonatal intensive care unit (NICU) of the West China Second Hospital of Sichuan University were enrolled in this study. In order to explore the characteristics of early microbiota on the gut-lung axis, we studied the correlation between intestinal and pharyngeal microbiota on day 1 and day 28 after birth in premature neonates. Microbial colonization in early life plays an important role in regulating intestinal and lung function. Objective: There are mutual influences between intestine and lung, that propose a concept of the gut-lung axis, but the mechanism is still unclear. 
2Key Laboratory of Birth Defects and Related Diseases of Women and Children (Sichuan University), Ministry of Education, West China Second University Hospital, Sichuan University, Chengdu, China.1Department of Pediatrics, West China Second University Hospital, Sichuan University, Chengdu, China.Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2.Sen Yang 1,2 †, Lina Qiao 1,2 †, Jing Shi 1, Liang Xie 2, Yang Liu 2, Ying Xiong 1 * and Hanmin Liu 1,2 * Zaneveld, Yilong Zhang, Qiyun Zhu, Rob Knight & J. van der Hooft, Fernando Vargas, Yoshiki Vázquez-Baeza, Emily Vogtmann, Max von Hippel, William Walters, Yunhu Wan, Mingxun Wang, Jonathan Warren, Kyle C. Torres, Pauline Trinh, Anupriya Tripathi, Peter J. Robeson, Patrick Rosenthal, Nicola Segata, Michael Shaffer, Arron Shiffer, Rashmi Sinha, Se Jin Song, John R. 
Peoples, Daniel Petras, Mary Lai Preuss, Elmar Pruesse, Lasse Buur Rasmussen, Adam Rivers, Michael S. Navas-Molina, Louis Felix Nothias, Stephanie B. Langille, Joslynn Lee, Ruth Ley, Yong-Xin Liu, Erikka Loftfield, Catherine Lozupone, Massoud Maher, Clarisse Marotz, Bryan D. Kelley, Dan Knights, Irina Koester, Tomasz Kosciolek, Jorden Kreps, Morgan G. Gibson, Antonio Gonzalez, Kestrel Gorlick, Jiarong Guo, Benjamin Hillmann, Susan Holmes, Hannes Holste, Curtis Huttenhower, Gavin A. Edwardson, Madeleine Ernst, Mehrbod Estaki, Jennifer Fouquier, Julia M. Cope, Ricardo Da Silva, Christian Diener, Pieter C. Callahan, Andrés Mauricio Caraballo-Rodríguez, John Chase, Emily K. Bisanz, Kyle Bittinger, Asker Brejnrod, Colin J. Alm, Manimozhiyan Arumugam, Francesco Asnicar, Yang Bai, Jordan E.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
